Genomics England, a leading provider of whole-genome sequencing diagnostics for the UK’s Genomic Medicine Service, aimed to scaleoperations to process 300,000 samples by 2025. To support this growth, they needed to transition from their internal workflow engine (Bertha) to a more scalable and efficient solution—Genie—built on Nextflow and the Sequera Platform.
Challenge
The migration required preserving the integrity of genomic diagnostics while ensuring automation, scalability, and efficiency. The transition needed to be seamless, with minimal disruption to existing workflows supporting cancer, rare disease, and population-scale studies.
Approach
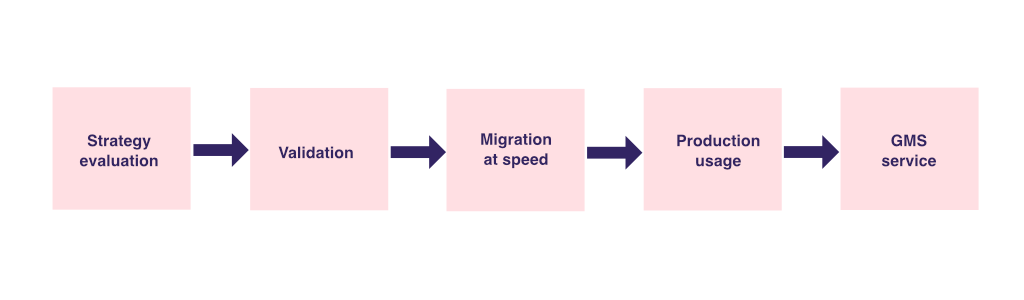
The project followed a structured migration strategy:
- A “Lift and Shift” approach was adopted to ensure a smooth transition while maintaining patient benefits.
- Agile methodology with bi-weekly sprints and continuous stakeholder feedback enabled iterative improvements.
- The Jasmine testing framework and containerized comparison tests ensured workflow reliability.
Results
- Regression testing time was reduced from days to minutes.
- Successful migration to Nextflow, enhancing automation and orchestration of bioinformatics workflows.
- Improved efficiency in processing a growing volume of sequencing data.
3 key programmes
8+ workflows
9+ bioinformatics components
- Cancer
- Rare disease
- Generation study
Along with its automation and orchestration
Conclusion
By leveraging a combination of off-the-shelf and customized bioinformatics solutions, Genomics England successfully modernized its genomic workflow infrastructure. The transition to Nextflow provided a scalable, high-performance environment, positioning the organization to meet its ambitious sequencing goals.
Expert Contribution
Reviewed by: Dr. Kamil Malisz, PhD
Role: Lead Workflows Developer and Solution Architect, AI‑Driven Drug Discovery
Expertise: Bioinformatics pipeline development, Nextflow workflow engineering, phage display and whole-genome sequencing, life sciences software engineering, project and team leadership.




