Metabolome 101 – What? Where? How?
What is a metabolome – in a few brief words.
Metabolomics is the study of metabolites, which are small organic compounds (usually weighing between 50 and 1500 daltons [Da]) that can be found within cells, biofluids, tissues, or whole organisms. They are typically involved in biological processes as intermediate or end products of the metabolism. The complete set of metabolites inside a biological system and their interactions at any given time point are known as the metabolome. As metabolites are continuously involved in multiple chemical reactions interacting with other molecules within and between biological systems and the environment, the metabolome itself is very dynamic and is dependent on multiple factors. Concentrations of metabolites may vary over time and that instability can be used to search for discrepancies, for instance, between healthy and the diseased individuals [3]. It may also be harnessed to identify interactions between a host’s metabolome and its microbiota.Methodologies used to study the metabolome
Metabolomics is a field that involves analyzing biological systems as the presence of particular metabolites and their concentration directly reflects underlying biochemical activity within cells or tissues [19]. There are a variety of methodologies used in metabolomics, but they all have their pros and cons and no tool is universally applicable to every analysis. To design a metabolomics study, one first has to decide on which approach to use: targeted or untargeted. The chosen approach will determine not only the design of the experiment, but also sample preparation and analytical techniques to be used. Untargeted methods are used to analyze samples by detecting and quantifying as many metabolites as possible, without any prior assumptions. On the other hand, the targeted approach is usually used to measure a particular group of metabolites to investigate the effects of a treatment or genetic modifications. Each of these approaches include multiple techniques to choose from when performing a metabolomic analysis. The most popular techniques which are used worldwide are Mass Spectrometry (MS) and Nuclear Magnetic Resonance (NMR) spectroscopy [5]. NMR is based on a physical phenomenon of nuclei of compound atoms that absorb and re-emit energy in the constant magnetic field created by a superconducting magnet. The characteristic resonance frequencies of the magnetic field are key features that allow for the prediction of the chemical structure of analyzed sample substances. The other technique, which is MS, is based on the mass-to-charge ratio . Unlike MS, NMR is quantitative and does not require extra steps during a sample’s preparation, such as the separation of a mixture (liquid or gas chromatography), and it does not destroy samples during analysis [20]. Unfortunately, the NMR sensitivity is significantly lower than that of MS, thus NMR alone may not be sufficient to obtain an in-depth understanding of samples [18]. At the same time, it is worth considering that if samples consist of metabolites that are not in mass spectral libraries, they might not be identified correctly by MS. Since stool samples are abundant with bacteria-borne metabolites (some of which are still to be thoroughly characterised), it’s beneficial not to rely on one method only and carefully plan the experiment in advance [1].Metabolome and microbiome combined: Multiomics data
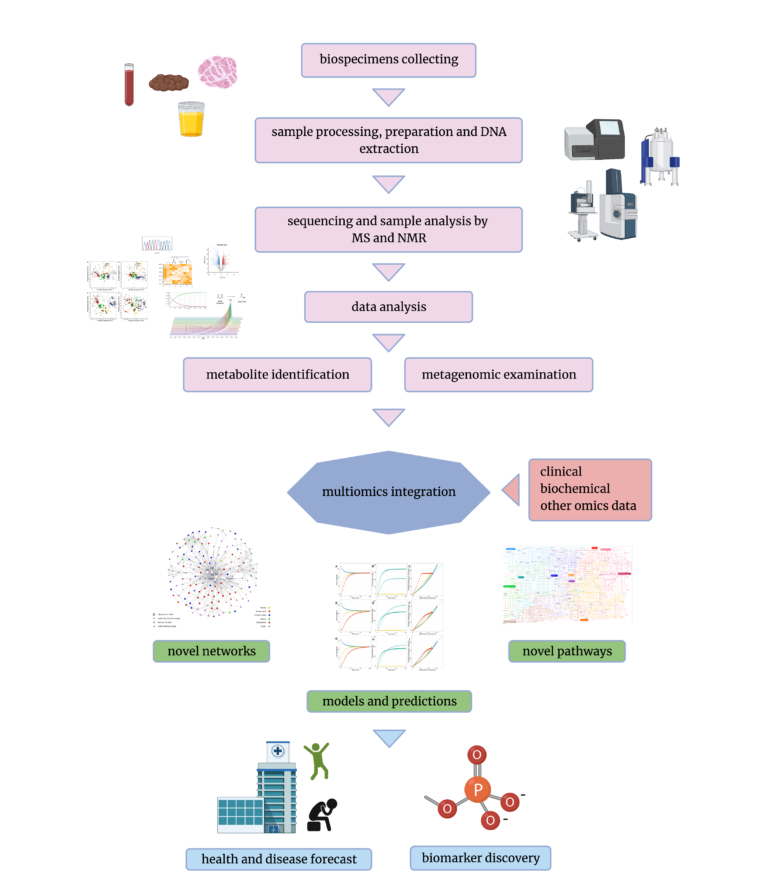
The most often used sample types in metabolomics are biofluids, such as stool samples, urine, cerebrospinal fluid or blood/serum, as they are the richest in metabolites. [4]. The presence or even a number of particular compounds in those can be an indicator of a disease, the effect of drugs or environmental factors, such as lifestyle or nutrition [10, 15, 17]. But what if we would like to see the bigger picture, not only the final product of the metabolism? Why not use the same sample for other purposes? This is where multiomics comes in – an approach that combines multiple types of data to achieve one goal. Currently, with the growing popularity of AI, especially deep learning, which is enabling the analysis of great volumes of data, the usage of combined systems is increasing [2, 11, 13]. After samples are collected from patients, they can be aliquoted and prepared for both sequencing (Whole Genome/ Exome Sequencing or Shotgun Metagenomics Sequencing) and metabolomics analysis (NMR and/ or MS) [12, 14]. Using the advantages of both allows for the discovery of underlying mechanisms that regulate interactions between gut microbiota and metabolites.Healthy/ Disease stages and corresponding metabolites (patient monitoring).
Detection of metabolites in blood or urine is commonly used for diagnostic purposes, as well as for monitoring a disease’s progression or a patient’s recovery. It can even help to understand whether a person is responding to the therapy or not. In the case of long-term treatment, the assessment of information on metabolite levels, and possibly other omic data, can be crucial for making informed decisions about a therapy’s (dis)continuation. In general, the more information the doctor/medical team has, the better they can tailor the treatment to patients’ needs.Where metabolomics is overlapping with the microbiome and how this helps us understand the bigger picture.
Humans are holobionts – complex ecosystems of bacteria and other microbial organisms interacting with each other and their hosts. Thus, the human metabolism should in fact be looked at from a wider perspective than just the processes that hosts are able to carry out themselves [6]. Adding information coming from gut metagenomes – pooled genomes from microorganisms present in one’s intestines and involved in food digestion – allows for a better understanding of the holobiont’s abilities to process particular compounds. Combining this knowledge with that of the metabolomics – meaning the actual metabolites present in the sample – finally allows for the unraveling of the network of dependencies between external dietary intake and internal nutrient metabolism [7, 8].Metabolite databases.
Since its formal date of birth in 1940, up until the present day, metabolomics has come a long way, and so have the collections of quantified metabolites. Information on them is deposited in multiple, mostly freely available, online databases. One of these is MetaCyc, which contains an encyclopedia of the metabolism, associated metabolites, enzymes and reactions, as well as a database of metabolic pathways from broad spectrum domains of life (selected bacteria, yeast, plants, human or model organisms like mice). Another database, HMDB – Human Metabolome Database [9], was designed to link chemical, clinical, and molecular biology/biochemistry data in one place. It contains a tremendous amount of information on common toxins, environmental pollutants, metabolic and disease pathways, as well as food components and microbial metabolites.An important bacterial sidestory.
Among records in HMDB one can find not only metabolites produced by humans but also some that are exclusively microbiota-derived. This may feel counterintuitive but is in fact well thought through as studies show they can significantly affect the host [16]. For instance, one of the most prominent members of this class of compounds are short chain fatty acids (SCFAs). Butyric acid (Butyrate) is a flagship SCFA and a metabolite, formed in the human colon by bacterial fermentation of carbohydrates. Butyrate is produced as the end-product of a pathway, solely present in obligate anaerobic bacteria, belonging to genus like Anaerostipes, Coprococcus, Eubacterium or Faecalibacterium. This bacterial metabolite has effects on cellular proliferation, apoptosis and differentiation. For more specific information on how SCFAs affect the host, see our previous posts “Short-chain fatty acids one step closer to the clinic” and “Microbiome mysteries: What happens in the gut does not stay in the gut”.Coming to conclusions: the inseparable bond between microbiome and metabolome
Integrating metagenomics with metabolomics and other omics could provide a multitude of invaluable information, synergistically enhancing the standard, one-dimensional, perspective. This approach to biospecimen analysis brings us to a place where nothing is wasted and the bigger picture is clearer. AI-powered multiomics is the future of tracking metabolic pathways potentially involved in patient health status and disease treatment. In general, multiomics integration holds promise in enabling the discovery of meaningful signatures, leading to biomarker discovery and the design of new therapeutics. At Ardigen, we are thrilled to work at the forefront of this exciting new field of study and hopefully contribute to unlocking the potential of personalized medicine in the near future.References:
- Sarma, S. J., Lei, Z., Rosenfeld, C. S., Ericsson, A., & Sumner, L. W. (2020). Nontargeted fecal metabolomics: an emerging tool to probe the role of the gut-microbiome in host-health. Bioanalysis. doi:10.4155/bio-2020-0010.
- O’Shea K, Misra BB. Software tools, databases and resources in metabolomics: updates from 2018 to 2019. Metabolomics. 2020 Mar 7;16(3):36. doi: 10.1007/s11306-020-01657-3. PMID: 32146531.
- Javdan B, Lopez JG, Chankhamjon P, Lee YJ, Hull R, Wu Q, Wang X, Chatterjee S, Donia MS. Personalized Mapping of Drug Metabolism by the Human Gut Microbiome. Cell. 2020 Jun 25;181(7):1661-1679.e22. doi: 10.1016/j.cell.2020.05.001. Epub 2020 Jun 10. PMID: 32526207.
- Lee-Sarwar KA, Lasky-Su J, Kelly RS, Litonjua AA, Weiss ST. Metabolome-Microbiome Crosstalk and Human Disease. Metabolites. 2020;10(5):181. Published 2020 May 1. doi:10.3390/metabo10050181
- Segers K, Declerck S, Mangelings D, Heyden YV, Eeckhaut AV. Analytical techniques for metabolomic studies: a review. Bioanalysis. 2019 Dec;11(24):2297-2318. doi: 10.4155/bio-2019-0014. PMID: 31845604.
- McBain, Andrew & O’Neill, Catherine & Amezquita, Alejandro & Price, Laura & Faust, Karoline & Tett, Adrian & Segata, Nicola & Swann, Jonathan & Smith, Adrian & Murphy, Barry & Hoptroff, Michael & James, Gordon & Reddy, Yugandhar & Dasgupta, Anindya & Ross, Tom & Chapple, Iain & Wade, William & Fernandez-Piquer, Judith. (2019). Consumer Safety Considerations of Skin and Oral Microbiome Perturbation. Clinical Microbiology Reviews. 32. 10.1128/CMR.00051-19.
- Tang ZZ, Chen G, Hong Q, et al. Multi-Omic Analysis of the Microbiome and Metabolome in Healthy Subjects Reveals Microbiome-Dependent Relationships Between Diet and Metabolites. Front Genet. 2019;10:454. Published 2019 May 17. doi:10.3389/fgene.2019.00454
- Harris EV, de Roode JC, Gerardo NM. Diet-microbiome-disease: Investigating diet’s influence on infectious disease resistance through alteration of the gut microbiome. PLoS Pathog. 2019 Oct 31;15(10):e1007891. doi: 10.1371/journal.ppat.1007891. PMID: 31671152; PMCID: PMC6822718.
- Wishart DS, Feunang YD, Marcu A, Guo AC, Liang K, Vázquez-Fresno R, Sajed T, Johnson D, Li C, Karu N, Sayeeda Z, Lo E, Assempour N, Berjanskii M, Singhal S, Arndt D, Liang Y, Badran H, Grant J, Serra-Cayuela A, Liu Y, Mandal R, Neveu V, Pon A, Knox C, Wilson M, Manach C, Scalbert A. HMDB 4.0: the human metabolome database for 2018. Nucleic Acids Res. 2018 Jan 4;46(D1):D608-D617. doi: 10.1093/nar/gkx1089. PMID: 29140435; PMCID: PMC5753273.
- Gopalakrishnan V, Helmink BA, Spencer CN, Reuben A, Wargo JA. The Influence of the Gut Microbiome on Cancer, Immunity, and Cancer Immunotherapy. Cancer Cell. 2018 Apr 9;33(4):570-580. doi: 10.1016/j.ccell.2018.03.015. PMID: 29634945; PMCID: PMC6529202.
- Lamichhane, Santosh & Sen, Partho & Dickens, Alex & Hyotylainen, Tuulia & Oresic, Matej. (2018). An Overview of Metabolomics Data Analysis: Current Tools and Future Perspectives. 10.1016/bs.coac.2018.07.001.
- Chen MX, Wang SY, Kuo CH, Tsai IL. Metabolome analysis for investigating host-gut microbiota interactions. J Formos Med Assoc. 2019 Mar;118 Suppl 1:S10-S22. doi: 10.1016/j.jfma.2018.09.007. Epub 2018 Sep 27. PMID: 30269936.
- Gardinassi, Luiz & Xia, Jianguo (Jeff) & Safo, Sandra & Li, Shuzhao. (2017). Bioinformatics Tools for the Interpretation of Metabolomics Data. Current Pharmacology Reports. 3. 1-10. 10.1007/s40495-017-0107-0.
- Bingol K, Brüschweiler R. Knowns and unknowns in metabolomics identified by multidimensional NMR and hybrid MS/NMR methods. Curr Opin Biotechnol. 2017 Feb;43:17-24. doi: 10.1016/j.copbio.2016.07.006. Epub 2016 Aug 20. PMID: 27552705; PMCID: PMC5305439.
- Ahmed I, Roy BC, Khan SA, Septer S, Umar S. Microbiome, Metabolome and Inflammatory Bowel Disease. Microorganisms. 2016 Jun 15;4(2):20. doi: 10.3390/microorganisms4020020. PMID: 27681914; PMCID: PMC5029486.
- Hernandez CJ, Guss JD, Luna M, Goldring SR. Links Between the Microbiome and Bone. J Bone Miner Res. 2016;31(9):1638-1646. doi:10.1002/jbmr.2887
- Shore SA, Cho Y. Obesity and Asthma: Microbiome-Metabolome Interactions. Am J Respir Cell Mol Biol. 2016;54(5):609-617. doi:10.1165/rcmb.2016-0052PS
- Emwas AH. The strengths and weaknesses of NMR spectroscopy and mass spectrometry with particular focus on metabolomics research. Methods Mol Biol. 2015;1277:161-93. doi: 10.1007/978-1-4939-2377-9_13. PMID: 25677154.
- Clish CB. Metabolomics: an emerging but powerful tool for precision medicine. Cold Spring Harb Mol Case Stud. 2015;1(1):a000588. doi:10.1101/mcs.a000588
- Schreier C, Kremer W, Huber F, et al. Reproducibility of NMR analysis of urine samples: impact of sample preparation, storage conditions, and animal health status. Biomed Res Int. 2013;2013:878374. doi:10.1155/2013/878374.